Watch a brief overview of what we do in the Roessler Group:
Our Research
In our group, we make use of chemical principles to understand fundamentally important reactions in biology. Reduction-oxidation (redox) reactions underpin innumerable chemical reactions and much of the chemistry of life. Many redox reactions proceed via radical intermediates and these are often located in mechanistically key locations (see our recent ). We investigate how oxidation-state changes govern respiration () and photosynthesis () and how nature has fine-tuned the redox properties of its many intricate molecular machines. Membranes play a fundamentally important role for many proteins and we are investigating the role of membranes on protein activity and function through spin labels and protein-intrinsic paramagnetic centres.
We are also developing film-electrochemical EPR (FE-EPR) as a new method to study the evolution of radicals during redox and catalytic reactions in real time. This enables detailed and previously impossible mechanistic understanding of both chemical and biochemical reactions. The catalytic reactions we investigate range from well-known long-established chemical transformations such as nitroxide-catalysed alcohol oxidation, to complex metal-containing enzymes that may prove to be novel targets for the development of new antimicrobials. Our is even finding applications in battery research.
Radicals often hold key mechanistic information - but trapping them in sufficient numbers for detailed EPR investigations can be a major challenge. We specialise in working with minute sample concentrations and volumes, and are seeking to make the impossible possible with our current collaborative grant .
(Protein) Film Electrochemistry-EPR
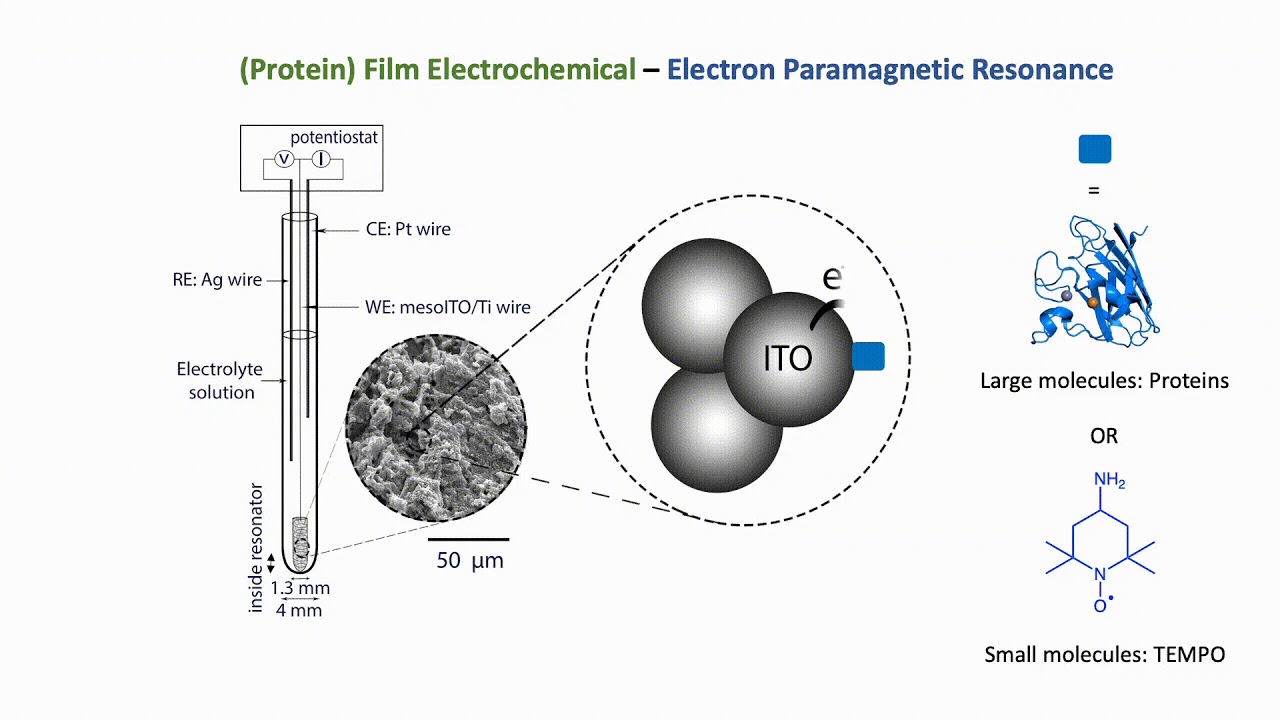
The combination of electron paramagnetic resonance (EPR) spectroscopy with protein film electrochemistry (PFE) provides a novel platform for the investigation of redox-active species including metalloproteins. The development of PFE-EPR enables the detection of paramagnetic species with direct and accurate potential control, providing new mechanistic insight to redox-based processes in biomolecules, including catalysis. Current EPR spectroelectrochemical methods to study proteins are based on generating radical species in solution and therefore limited by diffusion (precluding accurante potnetial control) and do not enable catalytic investivations. We exploit the advantages of PFE, i.e. 'wiring' electrons directly to the working electrode surface, to eliminate diffusion limitations.
PFE-EPR
Respiratory Complex I
NADH:ubiquinone oxidoreductase
Respiratory Complex I is the main entry point for electrons into the respiratory chain in higher organisms, but its mechanism is still not fully understood. In mitochondria, it oxidises NADH from sugars and fats, reduces ubiquinone, and pumps protons across the inner membrane. This process helps generate the proton motive force that drives ATP production.
Our work focuses on understanding how Complex I functions, particularly how energy from electron transfer is used to move protons. Because the enzyme is very large (around 1 MDa), many common techniques like NMR cannot be used. As such, we use EPR spectroscopy, which is blind to the many paired electrons in the enzyme, and instead detects unpaired electrons located at key functional sites in the enzyme. For example, iron–sulfur clusters ([2Fe–2S] and [4Fe–4S]) become EPR-visible when reduced.
We have developed a that allows us to precisely control the reduction state of small protein samples. Combined with pulse EPR, this approach lets us study the role of specific componants, such as the . Additionally, we are expanding our research and the techniques we have pioneered to look at the role Complex I plays in the heightened electron transport chain efficiency observed in glioblastoma - an aggressive brain cancer.
Photosynthetic Complex I
The NAD(P)H dehydrogenase-'like' complex (NDH)

We have teamed up with to investigate the fascinating molecular machine NDH, in many ways related to the mitochondrial enzyme (complex I, above), present in plants and cyanobacteria. Our goal is to unravel the mechanism of NDH, and ultimately to exploit this knowledge in order to increase stress tolerance in plants (in collaboration with , Rosario, Argentina). Increasing crop yields is an especially important challenge facing the developing world, but may affect us all one day given the ever rising world population. Research on NDH has thus far focused on genetic investigations and very little is known about the function of NDH, and no biophysical studies are reported. Using our experience with the mitochondrial enzyme, we are using EPR spectroscopy and protein film electrochemistry to investigate the co-factors in and the electron donors to NDH. Moreover, the knowledge gained through NDH is also likely relevant for understanding the mechanism of complex I.
See Dr Hanke's for more details on the biological context.
Artificial membrane systems: tailor-made tools to study membrane proteins

Biological membranes are mainly formed from bilayers of amphiphilic phospholipid molecules and harbour small hydrophobic components, such as vitamins, quinones and pigments. In addition, they can host a huge range of membrane proteins from small, peripherally bound proteins to integral multicomponent megaDalton complexes. The total protein content of a membrane increases with the complexity of the biochemical functions it helps to sustain.
The innate complexity of densely packed biological membranes makes targeted functional studies of individual membrane proteins in their native cellular environment challenging. To overcome this problem, mimetic systems ranging from membrane-monolayers to lipid vesicles (known as liposomes) have been employed. Such systems enable membrane proteins to be studied in relatively simple, well-defined environments while still being orientated and inserted inside a lipid environment needed for full activity.
The complex interaction patterns inside the used lipid bilayer structure participate significantly in the overall performance of the reconstituted protein of interest. Therefore, we (in collaboration with the Membrane Biophysics group) are interested in studying the membrane ordering and fluidity in the pure liposomes to fully unravel the interplay of biomimetic environment.
Funding